Abstract
Objective: Bipolar disorders have a strong genetic underpinning. Little is known about biological predispositions that convey vulnerability for the illness. We searched for biological vulnerability markers using proton magnetic resonance spectroscopy (MRS) in both affected and unaffected participants at high genetic risk for bipolar disorder.
Methods: We recruited high-risk participants aged 15–30 years from families in which multiple members were affected with bipolar disorder. Our primary sample included 14 affected and 15 unaffected relatives of probands with bipolar I disorder. Our extended sample comprised 19 affected and 21 unaffected participants with a family history of either bipolar I or bipolar II disorders. We matched both samples by age and sex with 31 control participants without a personal or family history of psychiatric disorders. We performed single voxel proton MRS at 1.5 T in bilateral dorsal and ventral medial prefrontal cortices with correction for grey matter proportion.
Results: We found comparable levels of choline, creatine, myo-inositol and N-acetylaspartate among the groups in both samples. There were no differences between regions of the medial prefrontal cortex or between hemispheres for any of the metabolites in any of the samples. The exclusion of 5 participants taking medication did not change our results.
Conclusion: Neurochemical changes in the medial prefrontal cortex that are measurable using proton MRS do not appear to be antecedent to the onset of mood disorders in genetically susceptible individuals.
Introduction
Bipolar disorder is a severe, recurrent psychiatric condition with heritability estimated at about 85%.1 The multifactorial etiology of the disorder is characterized by interactions between susceptibility genes and environmental factors. The presence of specific susceptibility alleles may lead to the expression of biological abnormalities, which develop before the onset of the illness and increase the risk of full clinical manifestation of the disorder.2 The identification of the biological predispositions for bipolar disorder is crucial for our understanding of the etiology, genetics, evolution, screening and early diagnosis of the illness.
Converging lines of evidence suggest that the ventromedial prefrontal and anterior cingulate cortices are involved in regulating mood.3 Changes in these regions have been detected in structural neuroimaging,4–6 neuropathologic postmortem, 7–9 functional neuroimaging4,10 and proton magnetic resonance spectroscopy (MRS) studies11–14 involving patients with bipolar disorder. The ventromedial prefrontal cortex is thus a suitable target in the search for biological abnormalities that convey vulnerability for mood disorders.
MRS is a noninvasive method of measuring concentrations of biochemicals in selected regions of the brain. Proton MRS investigations of children and adolescents with bipolar disorder reported a decreased N-acetylaspartate–to–creatine (NAA-to-Cr) ratio,15 an elevated level of myo-inositol (mI)12,13,16 and an increased glutamate-to-Cr ratio.17 Several studies found abnormalities in affected children with a family history of bipolar disorder — specifically a decreased NAA–Cr ratio18 and an elevated level of mI.16
To date, only 1 study has examined unaffected participants at genetic risk for bipolar disorder who did not yet have a diagnosable mood disorder. Gallelli and colleagues19 studied individuals who had bipolar parents and who also had bipolar disorder or subsyndromal symptoms of the disorder (e.g., moderate mood symptoms, diagnosis of attention-deficit hyperactivity disorder [ADHD]) and found no abnormalities of metabolite concentrations in the dorsolateral prefrontal cortex among experimental participants compared with control participants.
Despite the value of such an approach in the search for biological markers of vulnerability, we are not aware of other spectroscopic studies in unaffected relatives of patients with bipolar disorder. We thus used a high-risk design to investigate affected and unaffected participants at familial risk for mood disorders. We expected to find decreased levels of NAA and increased levels of choline (Ch) and mI in affected relatives of bipolar probands who were at an early stage of illness, with values for unaffected relatives falling between those of affected relatives and offspring of parents who did not have bipolar disorder (control group).
Methods
Participants
Families
We recruited the high-risk offspring according to the methods described elsewhere from families with multiple members with bipolar disorder.20 Briefly, we identified suitable families through adult probands with bipolar disorder who had participated in genetic studies and had been recruited from outpatient clinics at the Queen Elizabeth II Health Centre in Halifax, NS. In 1 case, the proband was a sibling, and in 4 cases the proband was a nephew or niece. In all other cases, the proband was a father or mother and the second parent was always unaffected. Each proband and affected parent completed a Schedule for Affective Disorders and Schizophrenia — Lifetime (SADS-L) interview21 conducted by 2 research psychiatrists blinded to the identity of the participant. An independent panel of senior clinical researchers, using all available clinical materials in a blind consensus fashion, reached final diagnoses according to Diagnostic and statistical manual of mental disorders, fourth edition (DSM-IV).22 To control for clinical heterogeneity, we performed primary analyses only in participants with a family history of bipolar I disorder (primary sample). Similar to previous studies,20,23 we also extended the sample by including participants with a family history of bipolar II disorders (extended sample — relatives of probands with bipolar I or bipolar II disorders). Probands with bipolar II disorder were similar to the participants with bipolar I disorder in that they experienced a low prevalence of comorbid conditions and an episodic course of illness. Bipolar II probands differed from bipolar I participants only in severity of mania. Family studies using similarly narrow diagnoses generally found bipolar II disorder to be a part of the same genetic spectrum as bipolar I disorder.24 Furthermore, with the exception of 2 studies, 25,26 most of the direct comparisons of main proton MRS metabolite concentrations (NAA, phosphocreatine plus Cr, glycerophosphocholine plus Ch, mI) did not find differences between participants with bipolar I and bipolar II disorders,27–30 or reported similar patterns of changes in participants with bipolar I and bipolar II disorders relative to control participants.25 One of the 2 positive studies used Diagnostic and statistical manual of mental disorders, third edition, Revised (DSM-III-R)31 definitions of bipolar II disorder,26 whereas the other one was likely confounded by differences in age and medication history between the groups.25 The lack of consistent differences between bipolar I andbipolar II disorders extends also to other neuroimaging modalities.32 A number of MRS investigations thus also studied both adults33–35 or children13,16 with bipolar I and bipolar II disorders.
High-risk offspring
Depending on their ages, a child/adolescent or adult psychiatrist interviewed the offspring using the Kiddie-SADS — Present and Lifetime (KSADS-PL)36 or SADS-L format. An independent panel of senior clinical researchers, in a blind consensus review, made the diagnoses according to DSM-IV and research diagnostic criteria. As part of the high-risk study, offspring were reassessed annually or at any time symptoms developed. The high-risk unaffected group comprised 21 offspring with no lifetime history of psychiatric disorders. The high-risk affected group comprised 19 offspring who met the criteria for a lifetime diagnosis of mood disorder and, in 1 case, psychosis not otherwise specified, which may be considered to be an antecedent of bipolar disorder. All high-risk affected participants were in remission at the time of scanning, as determined by a psychiatrist during an interview based on functioning at work or school and the absence of depressive or manic syndrome (Clinical Global Impression — Severity of Illness score of 0 or 1). Exclusion criteria included a history of closed head injury resulting in loss of consciousness, an untreated active medical illness (e.g., diabetes), an identified learning disability or diagnosis of ADHD, a substance-related disorder within the 6 months preceding our study, a lifetime history of substance dependence and a history of neurological disease.
Offspring of well parents (control participants)
We selected control participants by advertisement in similar geographic and sociodemographic areas as high-risk participants. The control group comprised 31 healthy offspring of well parents who were interviewed by a child/adolescent or adult psychiatrist in accordance with the KSADS-PL or SADS-L formats and deemed to be well upon blind consensus review. We selected the control participants to closely match high-risk participants in age, sex and parental education status. Exclusion criteria were the same as those in the high-risk groups, in addition to a personal or family history of psychiatric disorders.
Prior to conducting the assessments, all interviewers underwent extensive training consisting of participation in interviews, interviews under supervision and blind consensus diagnostic reviews. The quality of clinical data, which was based on both detailed baseline assessment and prospective follow-ups,20 was that required for genetic studies. We evaluated handedness by asking participants what hand they used for writing.
After complete description of the study to the participants, we obtained written informed consent. The Research Ethics Boards of the IWK Health Centre and Capital District Health Authority in Halifax, NS, approved the study.
MRI methods
MRI acquisition parameters
We performed all MR acquisitions with a 1.5 T Signa scanner (General Electric) and a standard quadrature head coil. We performed a T1-weighted spoiled gradient scan with the following parameters: flip angle = 40°, echo time (TE) = 5 ms, field of view (FOV) = 24 cm × 18 cm, matrix = 256 × 160 pixels, number of excitations (NEX) = 1, no interslice gap, 124 images 1.5 mm thick. We performed the scans at about the same time of day in all participants to minimize biological variability.
Proton MRS acquisition parameters
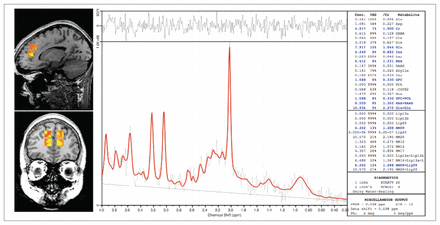
We performed 4 single-volume proton MRS acquisitions with a PROBE (proton brain exam) PRESS (point resolved spectroscopic sequence) and the whole gradient mode. The PROBE acquisition signal acquired the unsuppressed water and suppressed spectra from the same location. We used the unsuppressed water signal for eddy current compensation and for metabolite quantification. The parameters for the MRS acquisition were as follows: TE = 30 ms, repetition time (TR) = 2000 ms, 320 acquisitions, 2500 Hz spectral bandwidth, 2048 data points and duration 11.5 minutes. Blinded to participant status, we placed a 15 mm (anterior to posterior) × 12 × 12-mm spectroscopic region of interest (ROI) in the left and then right dorsal medial prefrontal cortex encompassing parts of the anterior cingulate, dorsal cingulate and medial parts of the dorsal prefrontal cortices (Brodmann areas 24c, 32 and 9). We aligned the anterior borders of the dorsal prefrontal cortex ROI with the anterior slice of the corpus callosum and the inferior borders with the upper limit of the horn of the lateral ventricles. We also placed a 13 × 13 × 13-mm ROI sequentially in the right and then left ventromedial prefrontal cortex, encompassing parts of the anterior cingulate and medial prefrontal cortices (Brodmann areas 32 and 10). We aligned the superior border below the inferior slice of the dorsal prefrontal cortex ROIs and placed the anterior border on the fifth slice anterior to the dorsal prefrontal cortex ROIs. We placed all 4 voxels as medially as possible while attempting to avoid pockets of cerebrospinal fluid (CSF) (ROIs are illustrated in Fig. 1). We manually placed outer-volume suppression bands close to each ROI as a further protection against the signals coming from the pericranial region or from orbital lipids.
An example of voxel placements on T1-weighted anatomical magnetic resonance scan (left) and proton magnetic resonance spectrum (right).
Tissue-type parcellation
The concentration of neurochemicals measurable with proton MRS varies substantially with the brain tissue type,37 which can lead to partial volume averaging effects in the estimated concentration of neurochemicals. To assess the partial volumes of grey matter, white matter and CSF within each ROI, we performed tissue-type segmentation using AFNI software. 38 We placed each ROI on the original T1-weighted images using the 3-dimensional coordinates that were defined during proton MRS acquisitions. We calculated the volumes of each type of tissue according to previously published criteria based on a histogram of tissue types for each selected volume of interest.39 These procedures for tissue-type parcellation have been extensively tested in our laboratory and have consistently shown a coefficient of reliability greater than 0.95 over several repeated assessments of the same ROI.
Spectral analysis
We quantified metabolite concentrations for NAA, Cr, Ch and mI using a linear combination model (LCModel) of in vitro spectra version 6.1, a commercially available automatic (user-independent) frequency domain fitting routine.40 The method employed a basis set of concentration-calibrated model spectra of individual metabolites to estimate absolute concentrations of similar brain metabolites from in vivo spectral data and corrected for residual eddy current effects and actual coil loading by using the transmitter reference amplitude. 41 The software provides concentration estimates with a confidence measurement (percentage of standard deviation [% SD]) reflecting maximum likelihood estimates and their uncertainties (Cramer–Rao lower bounds).40 The window of frequency domain data analyzed was left with the default settings: 0.2–4.0 ppm. An example of LCModel output is provided in Figure 1. We did not analyze the glutamine/glutamate peak because evaluation of this peak at 1.5 T is unreliable owing to J-coupling.
We determined the quality of the output data based on criteria outlined in the literature.42 We considered spectral profiles to have acceptable quality when the full width at half maximum was smaller than 0.08 (as measured by the LCModel program), when the signal-to-noise ratio was 6 or greater with randomly distributed noise and in the absence of artifacts in the spectral profile or of doublets in the peaks. Of the spectra that we judged to have overall acceptable quality, we retained only the estimated neurochemical concentrations with a % SD lower than 15%. Furthermore we excluded spectra with macromolecule contribution, as evidenced by distorted baseline of the spectral profile. Blinded to participant allocation, we excluded spectra based on the above-mentioned criteria. We reported the neurochemical quantifications in terms of institutional units. We used absolute values rather than ratios to other metabolites because this made the interpretation of results clearer and because the use of ratios was previously questioned. 43 A similar approach has been used previously in studies involving patients with bipolar disorder.12,44
At an echo delay of 30 ms, contribution from macromolecules will enter the spectrum. There is no consensus about how to deal with this unwanted contribution of macromolecules to spectral profiles. The LCModel tries to model most of them. Some authors prefer to completely eliminate the signal from macromolecules by deleting the first datapoints in the time domain of the free induction decay, where these large molecules predominantly resonate.15,35 In the absence of clear consensus, we decided to use both approaches, and we performed statistical analyses independently for each method.
To eliminate the macromolecules, we manually fitted the 3 main singlets of the short TE spectra in the time domain using the fitMAN analysis software.45 We converted the PROBE files to fitMAN format. We performed a lineshape correction using the quality deconvolution and eddy current correction (QUECC) technique to restore the Lorentzian line-shape. Prior to metabolite fitting, we removed the residual water peak using an operator-independent singular value decomposition fitting algorithm. We accomplished the fitting using a 3-peak constraint file (prior knowledge) and a 3-peak guess file. In the constraint file, we omitted 74 datapoints (delay time of fit = 29.6 ms) at the beginning of the free induction decay to minimize the influence of signals coming from the lesser amplitude peaks (the complex overlapping multiplets and the overlapping peaks of macromolecules). Such preprocessing has been shown to reliably and accurately quantify NAA, Cr and Ch. Using the guess file, we adjusted amplitude, frequency shift and line width for each spectrum visualizing the frequency domain to approximate the correct fit before generating the automated fitting algorithm. We fitted the unsuppressed water signal similarly, but we approximated the guess values visualizing the time domain. The compilation of estimated neurochemical levels included scaling using the unsuppressed water as an internal standard and an adjustment for the partial volumes of tissue types in each ROI. We did not perform an adjustment for potential between group differences in T1 and T2 relaxation times in the prefrontal cortex, therefore, we reported neurochemical levels in arbitrary units instead of in millimoles per litre.
To control for partial volume averaging effects, we covaried the metabolite results with grey matter proportion within a ROI. Cerebrospinal fluid contributions were negligible (about 1%). Because there were no differences in proportion of CSF among the groups, we did not deem it necessary to apply a scale factor for fractional content of CSF.
Statistical analyses
To control for clinical heterogeneity, we performed separate analyses of data from participants with a family history of bipolar I disorders (primary sample). To maximize power, we also analyzed an extended sample of participants with family history of either bipolar I or bipolar II disorders (extended sample). We used one-way analysis of variance (ANOVA) for comparisons of age among the 3 groups. We compared categorical demographic variables (e.g., sex, handedness) using the Pearson χ2 test. To be able to use all data that met our quality criteria, we used unbalanced repeated-measures models (module 5V in BMDP software, Statistical Solutions). To be conservative and not to enforce any a priori assumptions into the model, we used an unstructured covariance matrix. This method used maximum likelihoods to estimate parameters of a general class of models in which the expected values of the responses were described as arbitrary linear functions of a set of regression parameters and the within-participant covariances were modelled as functions of the set of unknown covariance parameters. These methods allowed for the use of incomplete observations. We sequentially used the concentrations of metabolites, proportions of grey matter and CSF as the dependent variables. We coded the ROI and laterality as within-subject factor, and we coded group (control, high-risk affected, high-risk unaffected) as between-subject factor. We used grey matter and age as covariates whenever they were significantly associated with the dependent variable. In some cases we recruited more than one participant per family. Presumably lower variance within individual families could lead to false-positive results; however, because none of our findings was positive, we did not deem it necessary to use mixed models to control for this.
We reported nominal p values, but in our interpretation of the results we corrected for multiple comparisons using a Bonferroni adjustment (4 comparisons for testing primary hypotheses). Controlling for 8 comparisons (4 metabolites in 2 samples) would be overly restrictive as the primary and extended samples overlapped.
We used the means and SDs from published papers to estimate Cohen’s d effect sizes (M1–M2/s pooled, where s pooled = √[(s12+ s22)/2]). We carried out an a priori power analysis for one-way ANOVA with 3 groups of participants. Effect sizes in previous positive MRS studies ranged from 0.7 to 1.246,47 for decreased levels of NAA, from 0.7 to 1.314,28 for increased levels of Ch and from 0.9 to 1.016,48 for increased levels of mI. Effect sizes in children ranged from 0.7 for decreased levels of NAA18 to 1.0 for increased levels of mI.12 To ensure 90% statistical power to detect an effect size of 0.7 at α = 0.05, we needed 17 participants per group. With 20 participants in each of the 3 groups (primary sample), we had 90% power to detect effect sizes of 0.6 at α = 0.05 and 80% power to detect effect size of 0.5 at α = 0.05. This was a conservative estimate of power because we had up to 4 measurements per participant, and we further increased our power in the extended sample.
We also calculated the percent difference that would be statistically significant. To make the most conservative estimate, we calculated these percent differences from the largest SD and smallest mean for each metabolite.
Results
Study population
Our primary sample included 14 affected and 15 unaffected participants with family history of bipolar I disorders. Of these, 24 had parents with bipolar I disorder and 5 had second-degree relatives with bipolar I disorder. Our extended sample included 19 affected and 21 unaffected participants with a family history of bipolar I or bipolar II disorders. Of these, 24 had parents with bipolar I disorder, 5 had second-degree relatives with bipolar I disorder, 9 had parents with bipolar II disorder and 2 had second-degree relatives with bipolar II disorder.
Participant characteristics are outlined in Table 1. Participants in both the primary and extended samples were closely matched by age, sex and parental education status to the 31 control participants who did not have a family or personal history of psychiatric disorders. We observed a smaller proportion of right-handed participants in the unaffected high-risk group in both the primary (χ22 = 5.97, p = 0.053) and the extended (χ22 = 4.63, p = 0.10) samples; however, we found no interaction between handedness and laterality of spectroscopic findings (concentrations in left v. right hemisphere) for any of the metabolites. Three affected participants in the primary group and 2 affected participants in the extended group used psychotropic medications at the time of scanning.
Demographics for the participants with a family history of bipolar I (primary sample) or bipolar I or II disorders (extended sample) and control participants without a personal or family history of mood disorders
Imaging
We found no differences in the proportion of CSF in ROIs among participants in both the primary and extended samples. We found no significant differences in the proportion of grey matter in ROIs in the primary or extended samples. Grey matter fraction was negatively correlated with Ch levels and positively correlated with creatine levels. Therefore we used grey matter as a covariate when analyzing differences in levels of Ch and Cr. We found no statistically significant association between age and any metabolite concentration.
In both primary and extended samples, the concentrations of Ch, Cr, mI and NAA that we obtained using the LCModel were comparable among the groups. Table 2 and Table 3 report the means and 95% confidence intervals (CIs) of metabolite concentrations in the primary and extended samples, respectively. Likewise, there were no differences among ROIs or between the left and right side and no interaction between group and laterality for any of the metabolites in either of the samples. We observed an interaction between the group and ROI for Ch in the extended sample. Unaffected participants and participants in the control group had similar Ch levels in the dorsal and ventral regions, whereas affected participants had lower Ch levels in the dorsal ROIs (χ22 = 6.02, p = 0.05) (Table 3). This interaction became nonsignificant when we controlled for the number of comparisons, and it was nonsignificant in the primary sample (χ22 = 4.43, p = 0.11) (Table 2). We found no interaction between the group and ROI for any other metabolite. Excluding the 5 participants who were taking medication at the time of scanning from the analyses changed our results for the interaction between group and ROI for Ch, which became nonsignificant. Our results did not change for any other metabolite. We found no differences between participants with a family history of bipolar I disorder compared with a family history of bipolar II disorder.
Concentrations of metabolites measured by proton magnetic resonance spectroscopy in the primary sample of affected and unaffected participants with a family history of bipolar I disorder and participants in the control group
Concentrations of metabolites measured by proton magnetic resonance spectroscopy in the extended sample of affected and unaffected participants with a family history of bipolar I or bipolar II disorders and participants in the control group
Overall, 77% of spectral profiles were available for our analyses of Ch, NAA and Cr levels, and 76% were available for our analyses of mI levels. We found no significant differences in the proportion of spectral profiles that did not meet one of our quality criteria for Ch or creatine levels among the groups. For NAA and mI, we excluded more spectral profiles for quality reasons among the unaffected participants relative to other groups. For NAA levels, the proportion of excluded spectra was 32% for unaffected participants, 17% for affected participants and 20% for participants in the control group (χ22 = 6.04, p = 0.05). For mI levels, the proportion of excluded spectra was 35% for unaffected participants, 18% for affected participants and 20% for participants in the control group (χ22 = 7.41, p = 0.03) (Table 2 and Table 3).
Results of manual fitting of spectra likewise showed no differences in concentrations of the 3 main metabolite peaks (NAA, Ch, Cr) among the groups, and they also showed no interaction between groups and ROIs for any metabolite.
An a priori power estimate showed that, with 20 participants in each of the 3 groups (primary sample), we had 90% power to detect effect sizes of 0.6 at α = 0.05. This allowed us to detect between-group differences of 3.18% for concentration of choline, 6.26% for concentration of NAA, 15.42% for concentration of mI and 4.63% for concentration of Cr as statistically significant.
Discussion
We found similar concentrations of metabolites measurable by proton MRS in unaffected and affected participants at high genetic risk for bipolar disorder and in control participants without a family or personal history of psychiatric disorders. The single study closest in design to our investigation was similar in terms of results. Gallelli and colleagues,19 in a study focusing on offspring of bipolar parents affected with or showing potential antecedents of bipolar disorder also found no differences between groups using proton MRS.
It is unlikely that the lack of significant differences between the groups in our study was a false-negative finding due to low statistical power. With 71 participants and up to 4 spectral profiles per participant, ours was one of the largest spectroscopic investigations of mood disorders. The largest, albeit nonsignificant, differences between groups that we observed were for Ch concentrations (p = 0.15). Based on a conservative estimate, we had more than 90% power to detect effect sizes smaller than those found in previous studies reporting abnormalities in Ch concentrations.11,28 In fact, our statistical power allowed us to detect statistically significant between-group differences of 3.18% for Ch, 6.26% for NAA, 15.42% for mI and 4.63% for Cr. Maybe with the exception of mI, it is unlikely that differences of the above-mentioned or lower extent have biological significance. Furthermore, as shown in Table 2 and Table 3, there was a marked overlap in 95% CIs for all pair-wise comparisons in all metabolites and all ROIs. Likewise, we observed no pattern of changes consistent with plausible biological explanations. We expected the greatest differences between patients and control participants, with unaffected relatives showing intermediate values between those of affected and control participants. Contrary to this, Ch levels were, in fact, largest among unaffected relatives of participants with bipolar disorder. Even if this represented a protective, compensatory effect, then affected participants should have had lower levels than participants in the control group. This was not the case for any ROI.
Clinical heterogeneity in the selection of parent probands and/or affected offspring might also account for our findings. Depression is most typically the first manifestation of an illness, even in patients who later receive diagnoses of bipolar disorder.49 Thus some of the currently unipolar participants at high risk for bipolar disorder may experience manic or hypomanic episodes in the future. In fact it has been estimated that about 70% of depressed first-degree relatives of bipolar probands also have bipolar disorder.50 To study early manifestations of bipolar disorders, the inclusion of unipolar participants with a family history of bipolar disorders is necessary. Likewise bipolar I and II disorders that are diagnosed conservatively may not be genetically independent. Family studies showed increased relative risk for bipolar II disorder among relatives of probands who have bipolar I disorder and vice versa.25 Exploratory analyses found comparable levels of neurochemicals between participants with a family history of bipolar I disorders and those with a family history of bipolar II disorders. This is consistent with other spectroscopic studies in which most direct comparisons have failed to find differences between participants with bipolar I and bipolar II disorders.34 Last but not least, separate analyses of participants with a family history of bipolar I disorders likewise revealed no differences between groups and no interactions. The remarkable consistency of our findings, regardless of our subgrouping of patients or methods of analysis of spectroscopic data, suggests that these are true-rather than false-negative findings. Overall, clinical heterogeneity did not play an important role in our data, especially because we also excluded participants who also had other psychiatric comorbid conditions, including alcohol or substance abuse and ADHD.
Our findings contrast those from several other spectroscopic studies involving patients with bipolar disorder. Potential confounding factors need to be considered when interpreting the existing literature. Some of the neurochemical abnormalities detectable by proton MRS are likely related to mood state,11,28,51,52 whereas others may be secondary to treatment with antidepressants, lithium, atypical antipsychotic medications or lamotrigine,37,44,51,53,54 or to the presence of comorbid conditions such as ADHD.55 Concentrations of metabolites differ between grey and white matter.37 In light of evidence for measurable volumetric abnormalities in patients with bipolar disorder, controlling for partial volume averaging effects is necessary.
Previous studies that detected abnormalities early in the course of illness included patients with current symptoms of mood disorders;13 a history of exposure to psychotropic medications, including stimulants, antipsychotics, antidepressants and mood stabilizers;15,18 and a high prevalence of comorbid conditions.16–18 Several studies did not adjust for the proportion of grey matter and CSF in the ROI.13,16,17 Notably, 5 of 6 studies reporting neurochemical abnormalities in the anterior cingulate cortex measurable by proton MRS investigated individuals who were symptomatic at the time of scanning,11–14,56 and depression ratings correlated positively with Ch-to-Cr ratios.14 Two of 3 studies that included euthymic patients found no abnormalities in this region.16,46 Thus it is possible that changes observed in previous studies of affected and often symptomatic patients developed during the course of illness or were dependent on treatment or mood state. We carefully controlled for previously identified confounding factors, including the effects of mood, medication, comorbid conditions, age and partial volume-averaging effects. Furthermore, we used stringent quality criteria for inclusion of spectral profiles to ensure high validity of the results.
Alternative interpretations of our results are that the early vulnerability markers detectable through proton MRS are more prominent in areas of the brain other than those targeted in our study, or that metabolites measurable through proton MRS do not capture the processes that convey vulnerability for mood disorders. It is also possible that the vulnerability biological markers are not static and that they need to be unmasked by external triggers such as administration of antidepressants, psychostimulants, reserpine, tryptophan depletion, sleep deprivation or others.
Limitations
There are several limitations to this study, including the cross-sectional design. A prospective design would have better allowed us to capture changes in neurochemistry related to the burden of illness. Some of the affected participants had unipolar depression, but depression at a young age in the offspring of parents with bipolar disorder is the most frequent early manifestation of bipolar disorder.20,49 We did not use a standardized instrument for assessing handedness. There was a trend for lower proportion of right-handed participants in the unaffected group. This difference was unlikely to bias results as right- versus left-handed participants did not differ in laterality of findings. In some cases we recruited more than one participant per family, which may have biased toward false-positive findings owing to presumably lower variance within families. Since none of the findings was positive, we did not use mixed models to control for this. We used 1.5 T magnet strength, which was a standard magnetic field strength when we initiated our study. Measurements of the main metabolites could be performed well at this magnetic field strength. Furthermore, most available MRS data on patients with bipolar disorder were obtained on 1.5 T magnets. We did not control for smoking status in the MRS analyses.
Conclusion
In summary, this well-powered study found no evidence of neurochemical abnormalities in medial prefrontal regions of the brain antecedent to the onset of mood disorders. Thus, abnormalities in the medial prefrontal cortex that are measurable using proton MRS do not seem to be harbingers for the onset of mood disorders. The lack of abnormalities measurable with proton MRS early in the course of mood disorders is optimistic from the clinical point of view because the previously reported neurochemical abnormalities in the medial prefrontal and anterior cingulate cortices may not represent vulnerability traits; rather, they may be related to mood state or the burden of illness and, as such, may be preventable.
Acknowledgements
Supported by NARSAD Young Investigator Award to Dr. Hajek, grants from the Department of Psychiatry, Dalhousie University and Capital District Health Authority, and from Canadian Institutes of Health Research.
Footnotes
Medical subject headings: magnetic resonance spectroscopy; bipolar disorders.
Competing interests: None declared for Drs. Hajek, Propper, Schmidt, Carrey, MacQueen and Alda and Mses. Bernier and Slaney. Dr. Duffy is a sponsored academic of AstraZeneca and has received travel assistance and grant funding for research unrelated to this article.
Contributors: Drs. Hajek, MacQueen and Alda and Ms. Bernier designed the study. Drs. Hajek, Propper, Schmidt, Carrey, Duffy and Alda and Mses. Bernier and Slaney acquired the data, which Drs. Hajek and Alda and Ms. Bernier analyzed. Drs. Hajek, Carrey and MacQueen and Ms. Bernier wrote the article. All authors reviewed the written article and approved its publication.
- Received August 24, 2007.
- Revision received November 2, 2007.
- Revision received January 14, 2008.
- Accepted January 14, 2008.